Download the DNAMAN v10.0.2.128 Review (Comprehensive Sequence Analysis for Molecular Biology) Software from this link…
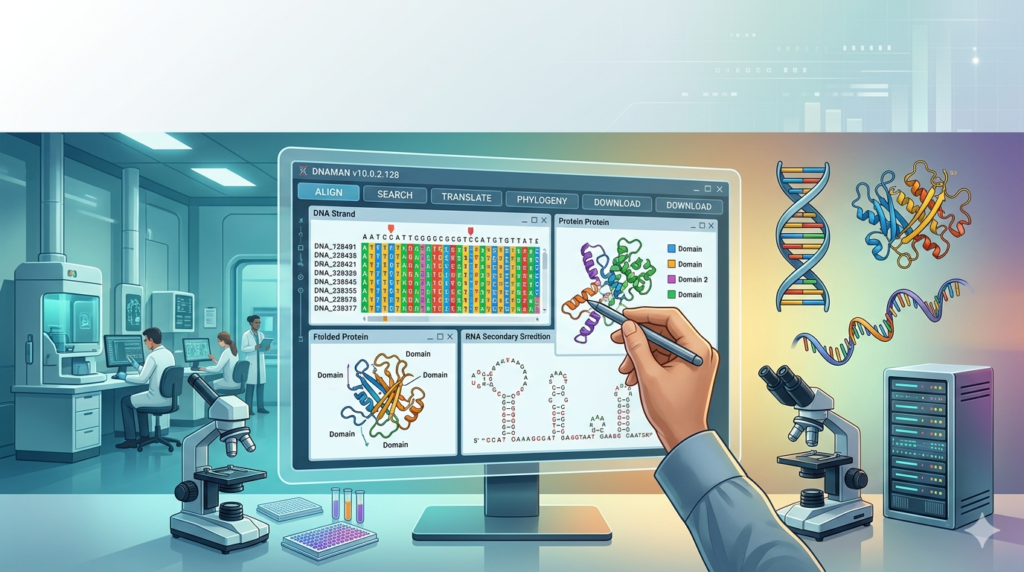
Overview of DNAMAN Software
Table of Contents
DNAMAN functions as an all-in-one solution for biological sequence analysis. Its primary purpose is to help scientists extract meaningful insights from genetic and protein data. Unlike single-purpose tools, DNAMAN combines sequence editing, alignment, restriction analysis, primer validation, and graphical visualization into a unified interface.
The software is commonly deployed in academic laboratories, pharmaceutical R&D, and agricultural biotechnology settings. Researchers rely on DNAMAN for gene expression studies, evolutionary biology, phylogenetic tree construction, and mutation detection. By offering validated analytical algorithms and reproducible results, DNAMAN supports both routine lab work and complex genomic research.
Key Features of DNAMAN v10.0.2.128
DNAMAN includes a robust set of features that address the full lifecycle of sequence analysis:
-
DNA and RNA Sequence Analysis – View, edit, and annotate nucleotide sequences. Generate reverse complements, translate DNA to protein, and analyze codon usage bias.
-
Protein Sequence Analysis – Compute molecular weight, theoretical pI, and amino acid composition. Predict secondary structures and identify functional motifs.
-
Multiple Sequence Alignment (MSA) – Align DNA, RNA, or protein sequences using proven algorithms. Identify conserved regions, insertions, deletions, and point mutations.
-
Restriction Enzyme Mapping – Simulate restriction digests, map cut sites, and predict fragment lengths. Essential for plasmid design and cloning strategy validation.
-
Primer Analysis and Design Validation – Assess primer melting temperature (Tm), GC content, hairpins, and dimer potential. Ensure PCR and sequencing reliability.
-
Graphical Visualization – Color-coded sequence views, alignment highlights, and editable restriction maps. Export high-quality figures for publications.
-
Phylogenetic Analysis – Construct evolutionary trees using neighbor-joining or maximum likelihood methods.
-
File Format Compatibility – Import and export GenBank, FASTA, EMBL, and other standard biological data formats.
What’s New in DNAMAN Version 10.0.2.128
The latest version introduces several improvements over earlier releases:
-
Enhanced alignment engine for faster processing of large sequence datasets (up to 10,000 sequences).
-
Improved graphical interface with customizable color schemes for annotation tracks.
-
Expanded restriction enzyme database with over 1,000 commercial and research enzymes.
-
Better integration with external databases for automated sequence annotation.
-
Optimized memory management for stable performance on Windows 10 and Windows 11 systems.
These updates make DNAMAN v10.0.2.128 more suitable for high-throughput sequencing workflows while maintaining accuracy for single-gene studies.
System Requirements
To run DNAMAN v10.0.2.128 effectively, your system should meet these specifications:
| Component | Minimum Requirement | Recommended |
|---|---|---|
| Operating System | Windows 10 (64-bit) | Windows 11 |
| Processor | Intel Core i3 or equivalent | Intel Core i5/i7 or AMD Ryzen 5+ |
| RAM | 4 GB | 8 GB or more |
| Storage | 500 MB free space | 1 GB SSD |
| Display | 1280 x 768 resolution | 1920 x 1080 or higher |
| Additional | Mouse and keyboard | Internet connection for updates |
Note: DNAMAN is designed primarily for Windows. For macOS or Linux users, virtualization or dual-boot setups may be required.
Installation Guide
Follow these steps to install DNAMAN v10.0.2.128 properly:
-
Download the installer – Obtain the official setup file from the developer’s website (Lynnon Biosoft).
-
Verify system compatibility – Ensure your Windows version meets the requirements.
-
Run as administrator – Right-click the installer and select “Run as administrator” to avoid permission issues.
-
Follow the setup wizard – Accept the license agreement and choose an installation directory (default:
C:\Program Files\DNAMAN). -
Select components – Choose full installation for all features or custom installation for specific tools.
-
Complete installation – Allow the installer to add necessary dependencies (e.g., Visual C++ Redistributables).
-
Activate license – Enter your valid license key or start the trial period. Do not use unauthorized activation methods.
-
Restart your computer – Recommended for proper registry and path configuration.
How to Use DNAMAN for Sequence Analysis
DNAMAN organizes its tools logically. Below is a typical workflow for analyzing a new gene sequence.
Step 1: Import or Create a Sequence
-
Use File > Import to load FASTA, GenBank, or text files.
-
Alternatively, manually enter or paste a sequence in the editor window.
Step 2: Edit and Annotate
-
Highlight coding regions, exons, or restriction sites.
-
Add comments and feature tags using the annotation toolbar.
Step 3: Perform Multiple Sequence Alignment
-
Select Analysis > Multiple Sequence Alignment.
-
Choose alignment parameters (gap penalty, matrix type).
-
Review the output for conserved motifs and variable positions.
Step 4: Restriction Mapping
-
Go to Tools > Restriction Analysis.
-
Select enzymes from the database or add custom recognition sites.
-
View the simulated gel pattern and fragment sizes.
Step 5: Primer Validation
-
Enter primer sequences under Tools > Primer Analysis.
-
Check Tm, GC%, and secondary structures.
-
Adjust primer design as needed for PCR or sequencing.
Step 6: Export Results
-
Save alignments as images (PNG, BMP) or text reports.
-
Export restriction maps for lab notebooks or publications.
Best Use Cases for DNAMAN
DNAMAN excels in several real-world research scenarios:
-
Academic Teaching – Demonstrating sequence alignment, translation, and phylogenetic concepts to students.
-
Cloning Project Planning – Simulating restriction digests and verifying insert orientation before wet-lab work.
-
Mutation Screening – Comparing patient or variant sequences against reference genomes to identify single nucleotide polymorphisms (SNPs).
-
Protein Engineering – Analyzing amino acid substitutions and their predicted effects on secondary structure.
-
Metagenomics – Aligning short reads from environmental samples to reference databases.
Advantages and Limitations
Advantages
-
All-in-one platform – Eliminates the need for multiple separate tools.
-
User-friendly graphical interface – Lower learning curve compared to command-line bioinformatics software.
-
Reproducible results – Consistent algorithms suitable for peer-reviewed research.
-
Offline functionality – No continuous internet connection required after installation.
-
Strong documentation – Built-in help and tutorial examples.
Limitations
-
Windows-only – No native macOS or Linux version.
-
Limited high-throughput support – May struggle with very large NGS datasets (use dedicated NGS pipelines instead).
-
Paid software – Free alternatives exist (e.g., BioEdit, UGENE, Jalview) though with fewer integrated features.
-
Updates require reinstallation – No automatic in-app updater.
Alternatives to DNAMAN
Depending on your needs, consider these alternatives:
| Software | Platform | Key Strengths | Best For |
|---|---|---|---|
| BioEdit | Windows | Free, lightweight sequence editor | Basic editing and alignment |
| UGENE | Win/Mac/Linux | Free, open-source, NGS support | Cross-platform workflows |
| Jalview | Cross-platform | Excellent alignment visualization | Multiple alignment analysis |
| Geneious Prime | Win/Mac | Advanced cloning and NGS tools | Professional biotech labs |
| SnapGene | Win/Mac | Superior cloning simulation | Molecular cloning design |
| MEGA | Cross-platform | Phylogenetic and evolutionary analysis | Tree building and evolution studies |
DNAMAN remains competitive for researchers who prefer a single, paid, Windows-native environment without the subscription cost of Geneious Prime.
Frequently Asked Questions (FAQ)
1. Is DNAMAN v10.0.2.128 compatible with Windows 11?
Yes, DNAMAN v10.0.2.128 runs stably on both Windows 10 and Windows 11 (64-bit editions).
2. Can DNAMAN analyze both DNA and protein sequences in the same project?
Absolutely. DNAMAN allows mixed-sequence projects, including translation between nucleotide and amino acid sequences and simultaneous alignment of both types.
3. Does DNAMAN support batch processing of multiple sequence files?
Yes. The software includes batch alignment and batch restriction analysis tools for processing dozens of sequences at once.
4. What file formats can DNAMAN import and export?
DNAMAN supports FASTA, GenBank, EMBL, PIR, SWISS-PROT, and plain text. Export options include sequence reports, alignment images, and restriction maps.
5. How does DNAMAN compare to free alignment tools like Clustal Omega?
DNAMAN offers a graphical interface and integrated features (restriction mapping, primer analysis) that free command-line tools lack. For pure alignment, free tools are sufficient; for a complete lab workflow, DNAMAN is more efficient.
6. Is there a trial version available before purchasing?
Yes. The developer (Lynnon Biosoft) offers a fully functional trial period, typically 30 days, with no feature restrictions.
7. Can I use DNAMAN for teaching in a university computer lab?
Yes. Educational and multi-user licenses are available. Contact Lynnon Biosoft directly for institutional pricing.
8. Does DNAMAN require an internet connection for core functions?
No. After installation and activation, all core analysis tools work offline. Only online help, updates, and optional database queries require internet access.
Final Thoughts
DNAMAN v10.0.2.128 remains a reliable, feature-rich platform for DNA, RNA, and protein sequence analysis. Its strength lies in combining multiple analytical tools—alignment, restriction mapping, primer validation, and visualization—into a single, coherent Windows application. For research laboratories that prioritize reproducibility, offline access, and a graphical interface, DNAMAN delivers consistent value.
While it is not designed for large-scale next-generation sequencing pipelines, it excels at routine molecular biology tasks, cloning project planning, and educational demonstrations. When compared to free alternatives, DNAMAN offers better integration and ease of use. When compared to premium tools like Geneious, it provides a more affordable perpetual license without subscription fees.
Premium Software Support Service
If you need professional help with software installation, setup, or technical configuration, our team is available to assist you.
Contact & Support
For quick assistance and latest updates, connect with us using the links below:
🔹 Direct Telegram Support
https://t.me/PlayoutKing
🔹 Official Telegram Updates Group
https://t.me/yourgroup
Service Policy
- Remote testing available through AnyDesk before confirmation.
• Verify the setup and performance before completing the order.
• Support available for single or multiple systems.
• Step-by-step guidance to ensure smooth installation and working environment.
Our goal is to provide reliable technical assistance so your software runs smoothly without interruptions.